Transcripts
The HRMDBQ prioritizes the MANE Select transcript by default. While this is the standard for variant curation, it is important to evaluate if the specific biology of a variant requires an alternative model.
Understanding MANE
Matched Annotation from NCBI and EMBL-EBI (MANE) transcripts represent exact matches between NCBI and Ensembl transcript models.
| Transcript Type | Description |
|---|---|
| MANE Select | A single transcript per locus, representative of the relevant biology. |
| MANE Plus Clinical | Additional transcripts for genes where "Pathogenic" (P) or "Likely Pathogenic" (LP) variants cannot be reported via the Select transcript alone. |
Example: PCSK9
In Blesa et al. (2009), researchers identified a variant affecting gene expression. Depending on the transcript model, the variant's location can be interpreted in multiple ways:
- Coding sequence
- 5' UTR sequence
- Upstream promoter sequence
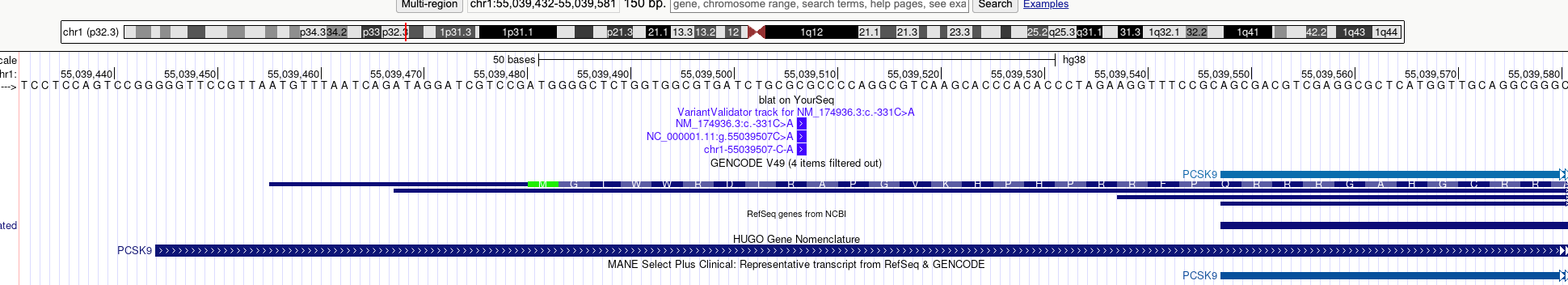
Figure 1: Promoter and 5' Untranslated Region of the F9 gene.
Variant Specification
The variant is located on Chromosome 1: NC_000001.11:g.55039507C>A. If we wanted to represent the variant as a 5' UTR sequence we would need to choose the corresponding transcript: NM_174936.3:c.-331C>A. However, the authors interpret the major effect as a promoter variant.
Evidence Summary
Luciferase reporter analysis indicated that the c.-332C>A variant caused a 2.5-fold increase in PCSK9 promoter activity relative to wild-type. Both patients with the mutation showed higher expression compared to samples with similar LDL-C levels.
Decision: We curate this as a Promoter variant because the preponderance of evidence suggests its effect is exerted via an alteration of promoter activity.
See also
- :material-microscope: Morales et al. (2022) – A joint NCBI and EMBL-EBI transcript set for clinical genomics and research. Nature.